Tandem MS (or MS/MS, MSn) is a technique to break down selected ions (precursor ions) into fragments (product ions). The fragments then reveal aspects of the chemical structure of the precursor ion.
The following scheme explains how Tandem MS works. Once samples are ionized (by ESI, MALDI, EI, etc.) to generate a mixture of ions, precursor ions of a specific mass-to-charge ratio (m/z) are selected (MS1) and then fragmented (MS2) to generate a product ions for detection. The selection-fragmentation-detection sequence can be further extended to the first-generation product ions. For example, selected product ions generated in MS2 can be further fragmented to produce another group of product ions (MS3) and so on.
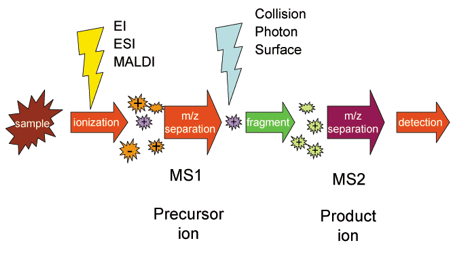
Since Tandem MS involves three distinct steps of selection-fragmentation-detection, the separation of these three steps can be realized in space or in time.
Typical Tandem MS in space instruments include QqQ, QTOF, and hybrid ion trap/FTMS, etc.
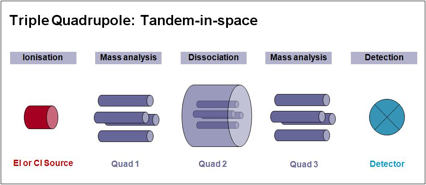
Three Quadrupoles (Quad 1, Quad 2, and Quad 3) are lined up in a row. Precursor ions are selected in Quad 1 and sent to Quad 2 for dissociation (fragmentation). The generated product ions are sent to Quad 3 for mass scanning.
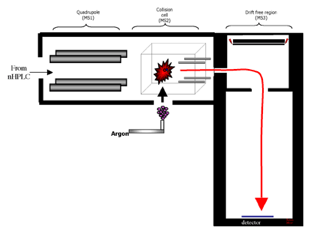
In the QTOF, precursor ions are selected in the Quadrupole and sent to the Collision Cell for fragmentation. The generated product ions are detected by time-of-flight (TOF) mass spectrometry.
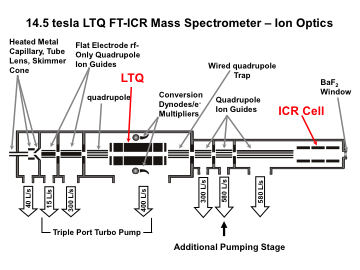
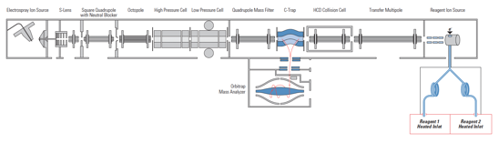
For the hybrid ion trap/FTMS (FT-ICR or Orbitrap) instruments, precursor ions are selected and fragmented in an external ion trap. The generated product ions can be detected either in the external trap (lower mass resolution, but faster) by or by FTMS (higher mass accuracy and resolution, but slower).
Typical Tandem-in-Time MS/MS instruments include ion trap and FT-ICR MS.
Peptides and oligosaccharides (including glycolipids) follow different systems of nomenclature for their fragment ions. Other classes of compounds, i.e. phospholipids, etc., do not yet have established nomenclature systems.
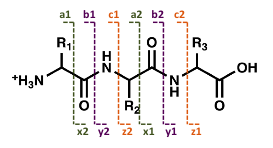
Fragments containing the N-terminus are labeled a, b, or c, depending on the site of the cleavage, whereas fragments containing the C-terminus are labeled x, y, or z. The numbers indicate the number of amino acid residues in the fragment ion.
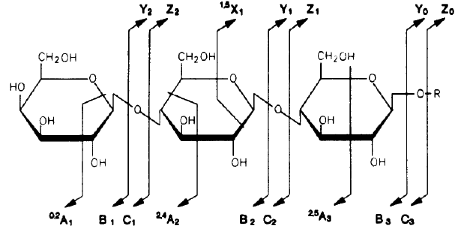
For oligosaccharides, fragments containing the reducing end (reducing end is on the right-hand side in the figure) are labeled x, y, or z, depending on the site of the cleavage, whereas fragments containing the other end are labeled a, b, or c. The numbers indicate the site of the sugar residue: y, z, b, and c ions are fragments due to glycosidic cleavages (cutting glycosidic bonds holding two adjacent sugar residues), whereas a and x ions result from cross-ring cleavage.
Precursor ions can be activated (with increased internal energy) in many different ways. Fragmentation patterns depend on how energy is transferred to the precursor ion, the amount of energy transferred, and how the transferred energy is internally distributed. Collision-induced dissociation and infrared multiphoton dissocition are "slow-heating" techniques that increase the Boltzmann temperature of the ion and thus preferentially cleave the weakest bonds to produce mainly b and y ions. These techniques are quite efficient for peptides, lipids and other relatively small chemical compounds, but may also remove protein post-translational modifications (e.g., phosphates and sugars). Electron capture dissociation and electron transfer dissociation mainly produce c and z ions while preserving post-translational modifications (PTMs). Thus, ECD and ETD are widely applied to proteins and peptides with labile PTMs. For oligosaccharides (including glycolipids), ECD/ETD can also generate cross-ring cleaved a and z ions, which are crucial for localization of glycosidic bonds.

Explore our magnet schedule to see what exciting research is happening on our stellar fleet of instruments right now.
B. J. Bythell, et al, Relative stability of peptide sequence ions generated by tandem mass spectrometry, Journal of the American Society for Mass Spectrometry 23(4), 644-654 (2012) Read online
For more information please contact Lissa Anderson.
Last modified on 16 March 2026